CCG develops a new method for mapping important sugars
The blood-thinning drug heparin is used worldwide. But the underlying structures and the related group of sugars, called heparan sulfates, have not been mapped in detail. Now, an international research group has developed a method for the detailed mapping of these types of structures. The method can contribute to uncovering important biological functions and thus to the development of new drugs. The study is led by researchers from the DNRF’s Copenhagen Center for Glycomics (CCG) at the University of Copenhagen and was recently published in the scientific journal Nature Communications.
There are many types of sugars, and they can be found everywhere both in nature and in humans, where they cover the surface of our cells. In particular, one type of sugar, polysaccharides, called GAGs (glycosaminoglycans), is found extensively in the human body. And in particular, one type of GAG, called heparan sulfate, is important for a variety of key biological processes, such as inflammatory conditions, neurodegeneration, and tumor metastases.
A type of heparan sulfate called heparin is one of the most widely used blood thinners in the world today. Researchers wish to map the underlying structures of heparin to have a clearer idea about its biological functions. However, only a few structures have been identified so far, but a new method has now been developed at the DNRF’s Copenhagen Center for Glycomics (CCG) at the University of Copenhagen to strengthen the mapping of these sugar structures.
“Determining the structures is a key question in the research about sugars. If we know the structure, we can determine what the cues are for specific biological functions and consider potential ways to exploit this in the development of therapeutics. This is hugely important and clinically relevant, as shown by the widely used anti-coagulant heparins, and the potential application of new heparin-based drugs for multiple diseases in the future,” said Rebecca Louise Miller, assistant professor at CCG and first author behind the study.
A virtual puzzle
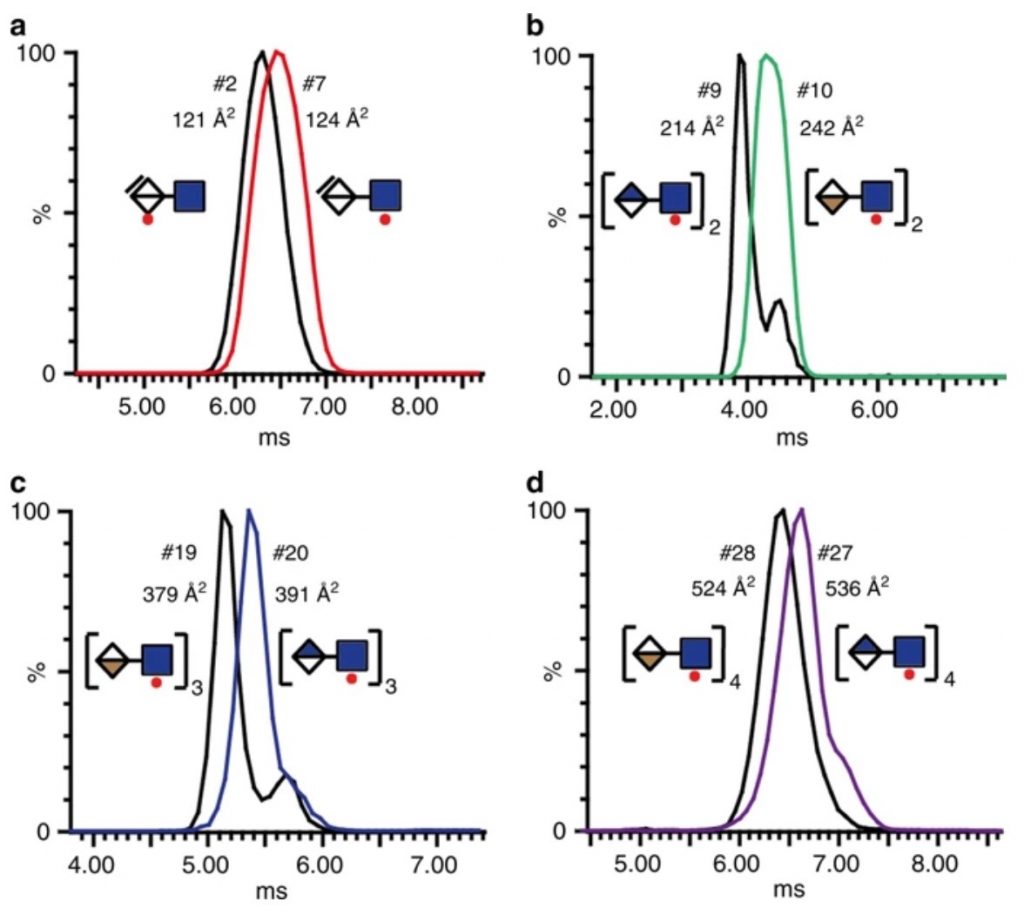
The new method is called “shotgun ion mobility mass spectrometry sequencing” (SIMMS2) and uses advanced mass spectrometry to divide the sugar structures into smaller pieces and take a sort of “fingerprints” of them for comparison to known standards. Subsequently, by virtually assembling the fingerprints in a large and complicated puzzle, Miller and the rest of the international research team can map, for the first time, extensive sequences of GAGs that are detailed enough to capture the signals that control important biological functions.
“The instrumentation behind this new method was invented by the company Waters Ltd in 2006 and is available to many pharmaceutical companies and researchers. This means that the method could be easily implemented and widely used for drug discovery by many research groups in a short period of time,” said Jeremy Turnbull, professor at CCG and co-author behind the study.
